Coronavirus Essay
What is Coronavirus?It is a group of closely related RNA viruses that infect animals and birds and spread diseases to humans. Both humans and other living species may experience mild to fatal respiratory tract infections due to the presence of this virus. Sometimes, people can have only colds, which are minor infections. However, more dangerous varieties can result in SARS, MERS, and other severe viruses, causing a pandemic that is still going strong worldwide. Due to the presence of such serious viruses, the human body becomes weak, and the immunity becomes so low that the body becomes vulnerable to other common diseases and has to face difficulty in fighting those diseases. They cause diarrhoea in cows, pigs and rats while also causing hepatitis and encephalomyelitis in pigs and cows. Coronaviruses have giant RNA virus genomes, ranging in length from 26 to 32 kilobases. Word OriginThe term "coronavirus" is derived from the Latin word corona, which means "crown" or "wreath". It is itself a derivation from the Greek word κορώνη korṓnē, which means "garland, wreath". When June Almeida and David Tyrrell first noticed and investigated human coronaviruses, they came up with the name. An unofficial group of virologists first used the term to describe the new family of viruses in the journal Nature in 1968. The viral spike peplomers, proteins on the virus surface, are responsible for this morphology. The International Committee on Taxonomy of Viruses, later known as the International Committee on Nomenclature of Viruses, recognised the scientific name coronavirus as a genus in 1971. The genus was split into four in 2009 as more new species were discovered: Alphacoronavirus, Betacoronavirus, Deltacoronavirus, and Gammacoronavirus. When did it start?The earliest reports of coronavirus infection emerged in the late 1920s when an acute respiratory illness of farmed hens arose in North America. The first thorough report describing a new respiratory disease in chickens in North Dakota was written in 1931 by Arthur Schalk and M.C. Hawn. The infection of young chickens was characterised by gasping and listlessness, with significant mortality rates of between 40 and 90%. Leland David Bushnell and Carl Alfred Brandly discovered the virus that caused the sickness in 1933. At the time, the virus was called the infectious bronchitis virus (IBV). In the 1960s, US and UK developed techniques to identify human coronaviruses. While working at the British Medical Research Council's Common Cold Unit, E.C. Kendall, Malcolm Bynoe, and David Tyrrell acquired the first samples of the distinctive common cold virus known as B814 in 1961. By serially transferring the novel virus through the organ culture of the human embryonic trachea, Tyrrell and Bynoe were able to cultivate it in 1965. Bertil Hoorn introduced the new cultivating technique to the laboratory. When the isolated virus was intranasally administered to volunteers, it produced a cold. It was rendered inactive by ether, demonstrating the presence of a lipid envelope. At the University of Chicago in 1962, John Procknow and Dorothy Hamre isolated a brand-new cold from medical students. The virus was identified as 229E after being separated and grown in kidney tissue culture. Like B814, the novel virus also gave volunteers a cold and was destroyed by ether. Tyrrell and Scottish virologist June Almeida investigated the structural differences between IBV, B814 and 229E in 1967 at St Thomas' Hospital in London. The three viruses could be distinguished from one another by their distinctive club-like spikes and general shape, thanks to electron microscopy. In the same year, a team of researchers at the National Institutes of Health used organ culture to isolate a second virus from this new family, designating one of the samples OC43 (organ culture). Under the electron microscope, the recent cold virus OC43 exhibited unique club-like spikes, similar to B814, 229E, and IBV. The newer cold viruses that resembled IBV were quickly determined to have morphological commonalities with the mouse hepatitis virus. Coronaviruses are the name of this new class of viruses because of their unique morphological characteristics. In the decades that followed, research on human coronavirus 229E and human coronavirus OC43 continued. SARS-CoV, HCV NL63, HCV HKU1, MERS-CoV, and SARS-CoV-2 are among the other human coronaviruses that have since been discovered. The structure of a CoronavirusThe structures of coronaviruses usually have distinctive surface projections and are significant, roughly spherical particles. Their sizes may range from typical diameters between 80 and 120 nm to extreme lengths between 50 and 200 nm. On average, the molecular mass is 40000 kDa (kilodalton). They are wrapped inside an envelope that contains numerous protein molecules. The nucleocapsid, membrane proteins, and lipid bilayer envelope shield the virus from the environment outside the host cell. The membrane (M), envelope (E), and spike (S) structural proteins are embedded in the lipid bilayer that makes up the viral envelope. In the lipid bilayer, the E: S: M molar ratio is roughly 1:20:300. The envelope usually has an 85 nm diameter. The lipid bilayer and the structural proteins E and M work together to shape and maintain the size of the viral envelope. The S proteins are also equally important as they are needed to interact with the host cells. 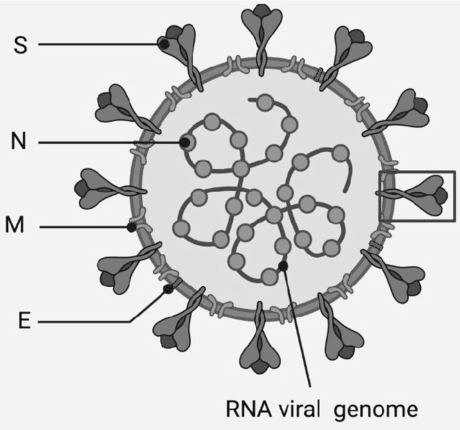
M proteinIt is a membrane protein that serves as the main structural component of the envelope and determines its general shape. It comprises layers that are 7.8 nm thick and range in thickness from 218 to 263 amino acid residues. It has three domains: an ectodomain at the short N-terminus, a triple-spanning transmembrane domain, and an endodomain at the C-terminus. The C-terminal domain creates a lattice-like structure that increases the envelope's extra thickness. The amino-terminal domain of a protein may contain N- or O-linked glycans, depending on the species. The viral lifecycle phases of assembly, budding, envelope formation, and pathogenicity depend on the M protein. E proteinIt is an auxiliary structural protein that differs significantly among species. A coronavirus particle contains only about 20 copies of the E protein molecule. They range in size from 8.4 to 12 kDa and contain 76 to 109 amino acids. They have two domains: a transmembrane domain and an extramembranous C-terminal domain. They are embedded in the lipid layer and form a five-molecular ion channel. They are almost entirely -helical, with a single-helical transmembrane domain. They are in charge of morphogenesis, intracellular trafficking, and virion assembly (budding). S proteinThe most distinctive characteristic of coronaviruses is their spikes. Typically, a coronavirus particle contains 74 spikes on its surface. Each spike is around 20 nm long and comprises the S protein. The S protein is made up of the subunits S1 and S2. The S protein is a fusion protein that encourages membrane fusion between the virus and the host cell and receptor binding. The S1 subunit, which includes a receptor-binding domain, makes up the spike's head. The stem, created by the S2 subunit, acts as an anchor for the spike in the viral envelope and facilitates fusion upon protease activation. The two subunits are exposed on the viral surface, and they remain noncovalently linked until they bind to the host cell membrane. NucleocapsidIt is enclosed inside the envelope, composed of many nucleocapsid (N) proteins, and continuously folds into the beads-on-a-string shape. It is connected to the positive-sense single-stranded RNA genome. The N protein is a phosphoprotein that contains three conserved domains and has a molecular mass of 43 to 50 kDa. Domains 1 and 2, typically enriched in arginines and lysines, make up most of the protein. Because there are more acidic amino acid residues than basic amino acid residues, domain 3 has a short carboxy-terminal end and a net negative charge. Transmission of VirusesCarriers who are infected can release viruses into the environment. Depending on the coronavirus species, they can spread from one host to another through an aerosol, a fomite, or a faecal-oral route. Coronaviruses infect the respiratory tract's epithelial cells. For instance, the SARS coronavirus attaches to the ACE2 receptor and enters the human lungs through an aerosol to infect the epithelial cells. OriginAlthough some studies state the most recent common ancestor of all coronaviruses to 55 million years or more, others claim it existed no earlier than 8000 BCE, suggesting long-term coevolution with bat and bird species. It has been estimated that the most recent common ancestor of the alphacoronavirus line, the betacoronavirus line, the gammacoronavirus line, and the delta coronavirus line occurred about 2400 BCE, 3300 BCE, 2800 BCE, and 3000 BCE, respectively. The origin of many human coronaviruses is in bats, and as evident as it is a prime natural reservoir for the coronavirus gene pool is found in bats and birds since they are warm-blooded flying animals. Coronaviruses have evolved and spread widely due to the vast number and variety of bat and bird species that may host viruses. Types of InfectionRisk factors differ significantly amongst coronaviruses. Some are comparatively mild, like the common cold. In contrast, others, like MERS-CoV, can kill more than 30% of infected individuals. Coronaviruses can bring on significant cold symptoms like fever and a painful throat from enlarged adenoids. Both primary and secondary bacterial pneumonia and bronchitis are brought on by coronaviruses (either direct viral or secondary bacterial bronchitis). There are six known species of coronaviruses, with one species (SARS) subdivided into two strains, for a total of seven strains of coronaviruses. Four coronaviruses generally cause mild symptoms, although they may have been more aggressive in the past:
The following three coronaviruses can cause severe symptoms:
Common ColdGenerally, rhinoviruses often cause the common cold. In contrast, coronaviruses are responsible for roughly 15% of cases of the common cold. The common cold's mild symptoms are caused by the human coronaviruses HCoV-OC43, HCV-HKU1, HCV-229E, and HCV-NL63, which are constantly circulating in the world's adult and paediatric populations. In temperate areas, the four mild coronaviruses are more common in the winter, and tropical regions do not have dominant seasons. 
Severe Acute Respiratory Syndrome (SARS)The World Health Organization (WHO) revealed in a news release that a new coronavirus found by many laboratories was the source of the severe acute respiratory syndrome (SARS) outbreak that began at rapid speed in Asia and spread to the rest of the world. SARS coronavirus (SARS-CoV) was the virus's official name. There were at least 774 fatalities out of over 8,000 infections across 29 nations and territories. The Middle East Respiratory Syndrome (MERS)In September 2012, a new coronavirus strain was found; it was first named Novel Coronavirus 2012 but is now known as Middle East Respiratory Syndrome Coronavirus (MERS-CoV). Soon after, the World Health Organization sent out a global alert. The virus did not appear to spread quickly from person to person, according to the WHO update from September 28th, 2012. On May 12th, 2013, the French Ministry of Social Affairs and Health confirmed a case of human-to-human transmission in France. Additionally, Tunisia's Ministry of Health reported instances of transmission from person to person. Two confirmed cases involved individuals who appeared to have contracted the illness from their deceased father, who became ill following travels to Saudi Arabia and Qatar. Despite this, it seemed that this virus also had trouble transferring from person to person in some cases because most infected people did not transmit it to others. There were 124 cases and 52 fatalities in Saudi Arabia as of October 30th, 2013. The virus was renamed Human Coronavirus-Erasmus Medical Centre after being sequenced by the Dutch Erasmus Medical Centre (HCV-EMC). Middle East Respiratory Syndrome Coronavirus is the full name of the virus (MERS-CoV). In May 2014, just two cases (both of which survived) were documented in the United States. In May 2015, a man who had visited the Middle East sought treatment at four hospitals in the Seoul area, resulting in a MERS-CoV pandemic. This resulted in one of the most severe MERS-CoV epidemics outside the Middle East. As of December 2019, laboratory tests had confirmed 2,468 cases of MERS-CoV infection, 851 of which were fatal, for a 34.5 per cent mortality rate. Coronavirus Disease in 2019 (COVID-19)In December 2019, a pneumonia outbreak was detected in Wuhan, China. On December 31st, 2019, the epidemic was linked to a new strain of coronavirus. The World Health Organization named it 2019-nCoV, but the International Committee on Taxonomy of Viruses termed it SARS-CoV-2. As of July 15th, 2022, the COVID-19 pandemic had resulted in at least 6,366,208 confirmed deaths and over 560,280,550 confirmed cases. The Wuhan strain was identified as a completely new Betacoronavirus strain from group 2B having a genetic resemblance to SARS-CoV of roughly 70%. Treatment and Cure
If an individual gets infected, vaccinating is the most excellent way to lower the chance of symptoms. Many vaccines against the coronavirus SARS-CoV-2 have been developed using various methods. Viral proteases, polymerases, and entrance proteins have also been identified as antiviral coronavirus targets. Drugs that target these proteins and the multiple stages of viral replication are being developed. However, in addition to vaccination, some precautions must be followed to avoid infection and potentially prevent the virus from spreading to others. They include using a mask when required, avoiding crowds, and keeping a physical distance.
Next TopicEssay on Water
|
 For Videos Join Our Youtube Channel: Join Now
For Videos Join Our Youtube Channel: Join Now
Feedback
- Send your Feedback to [email protected]
Help Others, Please Share









