Protein Synthesis | DNA Transcription, DNA Translation | Gene ExpressionProteins are building blocks of life. All cells in the human body contain proteins as it is the basic component of human body cells. Apart from water and fat, the human body is almost completely made of proteins. For example, protein is the main component of bones, organs, skin, nails, muscles, etc. The 80% portion of the muscles is made of proteins. Besides this, our body needs proteins to perform various vital functions such as transport of oxygen, to create muscles, regulate chemical reactions, etc. The proteins are made of small units or components called amino acids. We can say that a protein molecule is a long chain of amino acids. There are a total of 20 amino acids which can combine in different ways to build different types of proteins. What is gene expression?The process by which the information (genetic code) stored or encoded in a gene (a segment of DNA) is used to produce the protein from DNA or to produce other functional products like ribosomal RNA and transfer RNA from DNA is called gene expression. As protein is made when a gene is expressed, the gene expression and protein synthesis are often considered the same process. Let us know how the proteins are synthesized in our body! Protein Synthesis is a vital process that occurs continuously in all the cells of living organisms. As the name suggests, proteins are produced in this process. DNA contains the information (genetic code) that tells the cells how to make proteins. The flow of this information in a cell is from DNA to RNA to Protein. This flow of information can be divided into two parts DNA to RNA and RNA to Protein. The flow of information from DNA to RNA occurs through a process called DNA transcription. And, flow from RNA to proteins occurs through DNA translation. So, the process of Protein synthesis consists of two major processes DNA transcription and DNA translation. Let us understand each of them one by one! 1) What is DNA transcription (The first step of protein synthesis or gene expression)?DNA transcription: In this process, a molecule of mRNA is formed from one strand of DNA that acts as a template and is known as the template strand. Only a segment of one DNA strand takes part in transcription and is copied into mRNA. We can say that it is a process in which genetic information is copied from DNA segment (gene) of one strand of DNA into mRNA. The principle of complementarity governs the process of transcription. For example, for an adenine base of a DNA template, there will be a Uracil base on the newly formed mRNA as RNA contains Uracil instead of Thymine. The enzyme RNA polymerase uses the template strand to produce messenger RNA (mRNA). The RNA polymerase with the help of proteins called transcription factors binds to a specific sequence of bases within the gene (the segment of DNA). This sequence of bases is known as a promoter. The enzyme then pulls the two strands apart out of which one strand serves as a template strand (anti-sense strand or non-coding strand) which means it will be used to generate the mRNA. The other strand is called the non-template strand (sense strand or coding strand). It has the same sequence of bases as the mRNA to be formed. RNA polymerase does not need any primer, it simply starts the synthesis of mRNA at the start point and moves down along the DNA template (gene) and this process is called elongation. It synthesizes the mRNA as it moves reading the antisense strand from 3' to 5' end and generate the mRNA from 5' end and attaches RNA nucleotides to the 3' end as it moves forward. Here, the RNA is made of a ribose sugar and it has Uracil instead of Thymine. Once the RNA polymerase reaches at the end of the DNA template, the termination occurs. The enzyme RNA polymerase detaches itself from the template and the DNA returns to its original state or shape after producing an mRNA through transcription. What is transcription unit?It is the DNA segment that is located between the site of initiation and termination of transcription. We can say that it is the segment of DNA that is transcribed into mRNA along with the sequences required for its transcription. So, it generally consists of all regions that take part in DNA transcription such as a promoter, structural gene (an RNA-coding sequence) and a terminator. The structural gene contains the template strand and non-template (coding) strand. The components of the transcription unit are described below:
This process of transcription is polycistronic in prokaryotes which means it occurs at more than one locations on DNA simultaneously. Whereas, it is monocistronic in eukaryotes which means it (DNA transcription) occurs at one location when it gets completed then it may occur at other location. The process of transcription can be divided into three steps that include initiation, elongation and termination. i) Initiation: It occurs at the point or location on the structural gene where transcription starts. RNA polymerase enzyme identifies the place of initiation which is called the promoter region. Then RNA polymerase binds to the promoter and initiates the process of transcription, this is called initiation. The promoter region contains a specific sequence of bases, e.g. TATAAAA in eukaryotes. The sigma factor finds the sequence TATAAAA to identify the promoter and starts initiation from there. In eukaryotes, the sequence of TATAAAA is called TATA box. The promoter region is generally present before the transcription starting site of the structural gene. 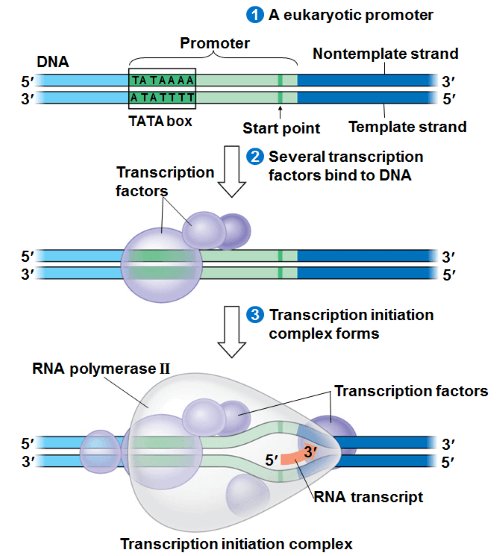
It is the sigma factors or transcription factors (a type of proteins) that help RNA polymerase to bind to the promoter region. The transcription factors start arriving and gathering at the promoter region that forms a transcription initiation complex. Now, this complex attracts RNA polymerase to the promoter region and tells RNA polymerase the location on the structural gene beside the promoter region from where it is supposed to start transcription. So, RNA polymerase with the help of transcription factors binds to the promoter region. Now, RNA polymerase starts unwinding the templates. The open strand looks like a bubble so it is called the transcription bubble. ii) Elongation: Once the strands start unwinding, the RNA polymerase starts the process of elongation. The helix starts opening and space is created between template-strand and coding-strand, and the mRNA starts getting elongated as the free nucleotides keep adding to the mRNA. 
After elongation starts the sigma factor gets separated from the core enzymes RNA polymerase and the core enzyme moves along the anti-sense strand (template strand) till it reaches terminator site. DNA ligase helps in the re-joining of strands by making pairs of nucleotides or bases. iii) Termination: It occurs when RNA polymerase enzyme reaches at terminator site and separates itself from DNA template. There is a rho factor to carry out the termination. This factor recognises the termination site and allows the polymerase to stop the process and to detach itself from the template strand. This step is shown below along with the first two steps. 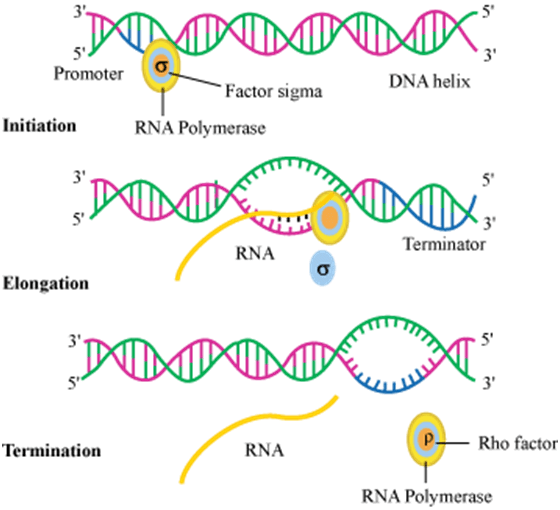
Difference between prokaryotic transcription and eukaryotic transcriptionAlthough initial steps are the same in both the types of transcription, there are lots of differences between them.
2) What is DNA Translation (The second or last step of protein synthesis or gene expression)?DNA Translation: The process of the formation of proteins from mRNA is called DNA translation. This process requires ribosomes, amino acids, mRNA, tRNA, and aminoacyl tRNA synthetase (an enzyme). Translation takes place in the cytoplasm at the ribosome, which is generally located on the rough endoplasmic reticulum. The ribosome is made of two units, a smaller subunit and a bigger subunit. The mRNA formed in transcription arrives at the ribosome and sits in-between the two subunits. The mRNA, as the name suggests, it contains a message or information for the formation of proteins. This information is present in the form of codons in the mRNA. So, the mRNA functions as a template for the formation of the polypeptide chain or protein. The sequence of three bases (codon) on m RNA code a specific amino acid. For example, AUG code for methionine. There is a molecule called tRNA that helps an amino acid finds its way to move to the corresponding codon on mRNA to become a part of the polypeptide chain. The transfer RNA (tRNA) transfer the amino acids to their respective codons. Translation occurs from 5' to 3' direction. How is tRNA loaded with amino acid?Let us understand in different steps, which are as follows: i) Activation of amino acids: First, the amino acid is activated. Amino acid and ATP react in the presence of Amino acyl t-RNA synthetase and produce Amino acyl AMP enzyme complex and two molecules of Phosphate. There are 20 types of amino acids that take part in protein synthesis.
Amino acid + ATP ---Amino acyl t-RNA synthetase----→Amino acyl AMP + PP
The enzyme (Aminoacyl tRNA synthetase) binds the amino acid to the tRNA. It always attaches an appropriate amino acid to the tRNA, for example, it will always attach methionine to a tRNA with UAC anticodon. UAC is complementary to AUG that codes for methionine. ii) Charging of t-RNA or loading of t-RNA: In this step, a specific activated amino acid is recognized by its specific t-RNA. It means a specific t-RNA is attached to a specific amino acid. The amino acid is attached to the 'Amino acid attachment site' of the t-RNA that forms amino acyl t-RNA complex and releases AMP and enzyme.
Amino acyl-enzyme complex + t-RNA ----→ Amino acyl t-RNA complex + AMP + enzyme
Amino acyl tRNA, which is a complex of amino acid and t-RNA, is called charged or loaded t-RNA. See the following image to understand the above steps. 
How transfer RNA knows which amino acid it has to place in the chain at which particular location?The tRNA also has a specific region that contains a specific sequence of bases that is called the anticodon. It is called anticodon as its sequence of bases is complementary to the base sequence (codons) on mRNA. For example, for AUG codon of mRNA, there will be a tRNA with UAC anticodon. Thus, a complementary base sequence or anticodon on tRNA helps it to attach to the complementary base sequence on mRNA. A specific amino acid based on the anticodon of tRNA is attached to the tRNA. The tRNA reads the codon on mRNA and attaches its anticodon with a complementary codon on mRNA. Thus, a tRNA is attached to the mRNA along with its amino acid or we can say that amino acid is transferred to the site of protein synthesis by tRNA. Similarly, another tRNA transfers another amino acid to the ribosome where bonds are formed between amino acids. The loaded tRNA arrives on the large subunit of the ribosome. There are three sites on the large subunit of the ribosome, which are End site (E-site), Peptidyl site (P-site), and Aminoacyl site (A-site). The first tRNA loaded with the amino acid arrives at P-site and the new tRNAs are landed at Aminoacyl site. At peptide site, the bonds are formed between amino acids. At the exit site, the tRNA leaves the ribosomes after transferring the amino acid. The translation also occurs in three steps that are initiation, elongation and termination. i) Initiation: As the name suggests, in this step, the process of translation starts. It occurs when the mRNA is attached to the small subunit of the ribosome and the reading of mRNA starts. There is an AUG codon on mRNA. It is a 'start codon'. So, translation starts from this codon. The large subunit of ribosome also joins the complex formed by the small subunit of ribosome, tRNA and mRNA to form the translation initiation complex. 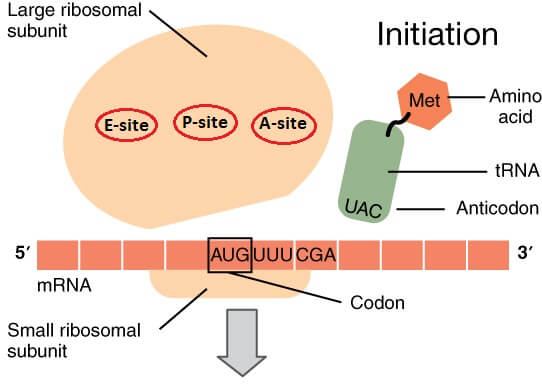
The tRNA with complementary codon UAC moves towards mRNA and attaches itself by pairing its anticodon UAC to AUG codon on mRNA. The first tRNA binds at P-site and rest of the incoming tRNA are loaded on Aminoacyl (A) site on the large subunit of the ribosome. In eukaryotes, there are 10 initiation factors that include eIF1, eIF2, eIF3, eIF4A, eIF4B, eIF4C, eIF4D, eIF4F, eIF5, eIF6. Whereas, in prokaryotes, there are three initiation factors that include IF1, IF2 and IF3. The initiation factors guide the tRNA about the location on mRNA at which initiation is supposed to occur. In eukaryotes, the binding site or smaller subunit where mRNA binds is recognized by "7mG cap". Whereas, in prokaryotes, like bacteria, there is a Shine-Dalgarno (SD) sequence (GGAGG) present on mRNA just before the start codon AUG. This sequence helps the mRNA to identify and binding with the small subunit of the ribosome. ii) Elongation: As the name suggests, in this step, elongation of the polypeptide chain starts as tRNAs keep bringing amino acids one by one. The new tRNA comes with new amino acid and gets attached at A-site of ribosome. Peptidyl transferase forms the peptide bond between two amino acids. Whereas, the translocase enzyme helps in the movement of the ribosome. The energy for the movement of the ribosome is provided by the GTP (Guanosine Triphosphate). There are two types of elongation factors in eukaryotes uEF1 and uEF2. Whereas, in prokaryotes, the elongation factors are of three types which are EF-Tu, EF-Ts, EF-G. 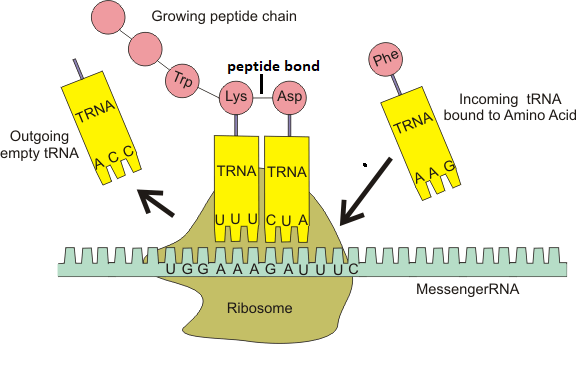
A peptide bond is formed between arriving amino acid and growing chain. The ribosome keeps moving forward along the mRNA as the elongation goes on. As the ribosome moves one codon position, the tRNA from A-site comes at P-site and again when ribosome moves one codon position, the rRNA from P-site comes at E-site (Exit site) and exit the ribosome from exit site and the chain gets ready to receive the next amino acid. We can say that as soon as the A site is vacated due to the movement of the ribosome and a new tRNA brings a new amino acid and occupies the A site. iii) Termination. As the name suggests, in this step, the translation stops. There are three types of stop codons if any one of them is present on mRNA and appears during elongation translation will stop. We can say that when ribosome is sliding over mRNA, any stop codon (UAA, UAG, UGA) is available at A site of ribosome then the process of translation terminates. In eukaryotes, there is only one release factor (eRF1) that helps the last tRNA to detach itself from the polypeptide chain. Whereas, there are three releasing factors (RF1, RF2 and RF3) in prokaryotes that along with GTP help the last tRNA in detaching itself from the polypeptide chain. 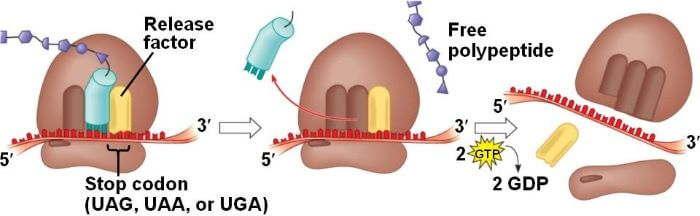
At this point, a tRNA which is having a release factor instead of an amino acid is loaded at the aminoacyl site. So, a new amino acid will not be added to the chain and the chain detaches itself from the tRNA and tRNA exits from the exit site and newly formed polypeptide chain will also leave the ribosome and thus termination will be completed. Difference between DNA transcription and DNA translation: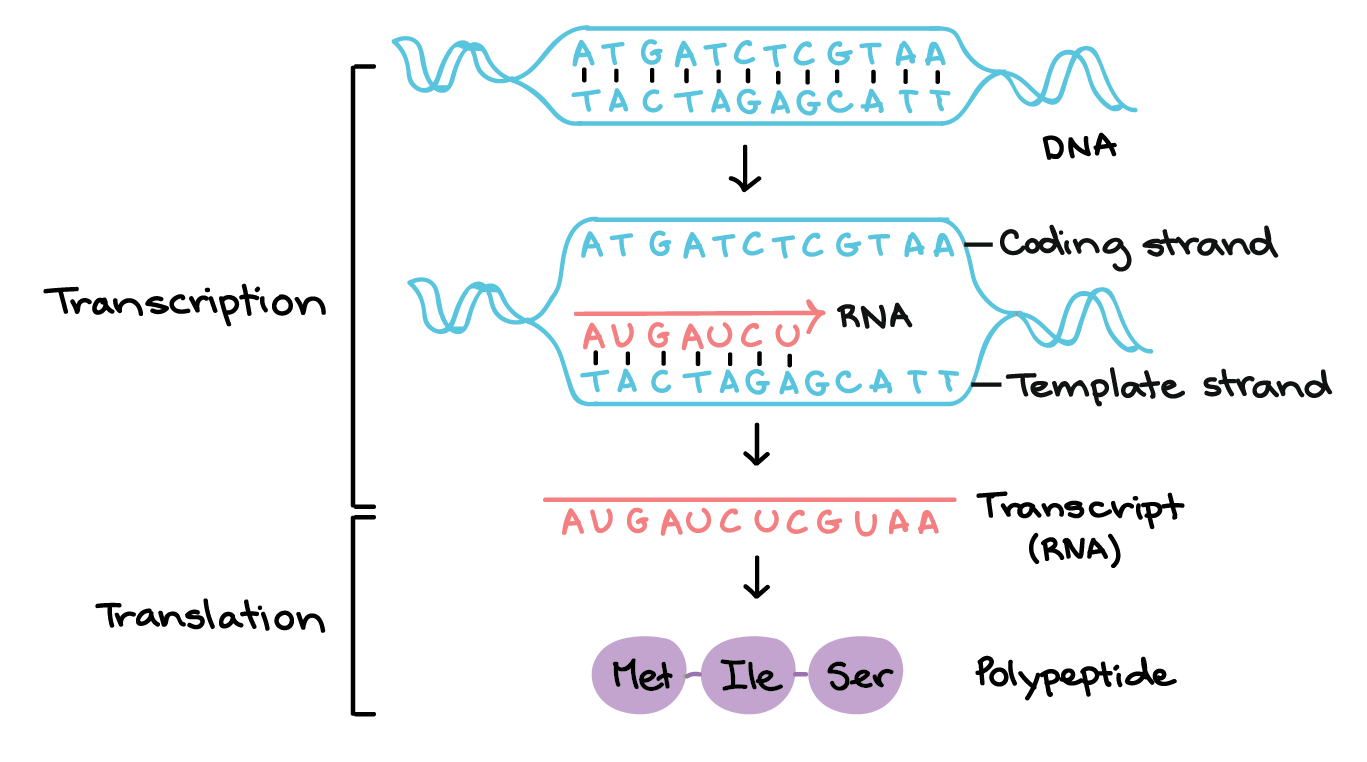
Next TopicAlgae
|
 For Videos Join Our Youtube Channel: Join Now
For Videos Join Our Youtube Channel: Join Now
Feedback
- Send your Feedback to [email protected]
Help Others, Please Share










