Probabilistic Model in Machine LearningA Probabilistic model in machine learning is a mathematical representation of a real-world process that incorporates uncertain or random variables. The goal of probabilistic modeling is to estimate the probabilities of the possible outcomes of a system based on data or prior knowledge. Probabilistic models are used in a variety of machine learning tasks such as classification, regression, clustering, and dimensionality reduction. Some popular probabilistic models include:
Probabilistic models allow for the expression of uncertainty, making them particularly well-suited for real-world applications where data is often noisy or incomplete. Additionally, these models can often be updated as new data becomes available, which is useful in many dynamic and evolving systems. For better understanding, we will implement the probabilistic model on the OSIC Pulmonary Fibrosis problem on the kaggle. Problem Statement: "In this competition, you'll predict a patient's severity of decline in lung function based on a CT scan of their lungs. You'll determine lung function based on output from a spirometer, which measures the volume of air inhaled and exhaled. The challenge is to use machine learning techniques to make a prediction with the image, metadata, and baseline FVC as input." Importing LibrariesEDALet's see this decline in lung function for three different patients. Output: 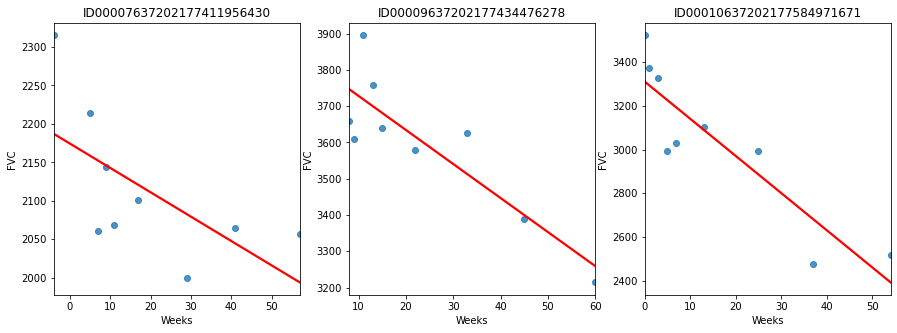
It is clearly obvious that lung capacity is declining. Yet as we can see, they vary greatly from case to patient. Postulate the modelIt's time to become imaginative. This tabular dataset might be modeled in a variety of ways. Here are a few tools we might employ:
We will start by attempting the most basic model, a linear regression, as we are still learning. We will, however, get a little more sophisticated. Following are our presumptions:
Our model is represented by the Bayesian network shown below: 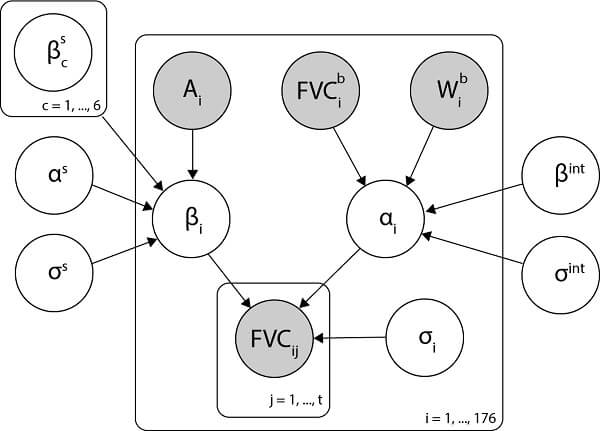
The logic behind this Model:
Mathematically Model Specification: 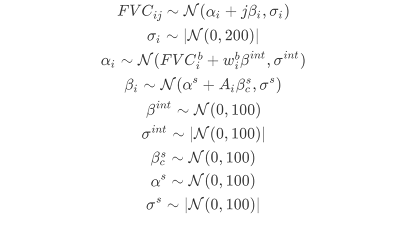
Simple Data Pre-processingOutput: 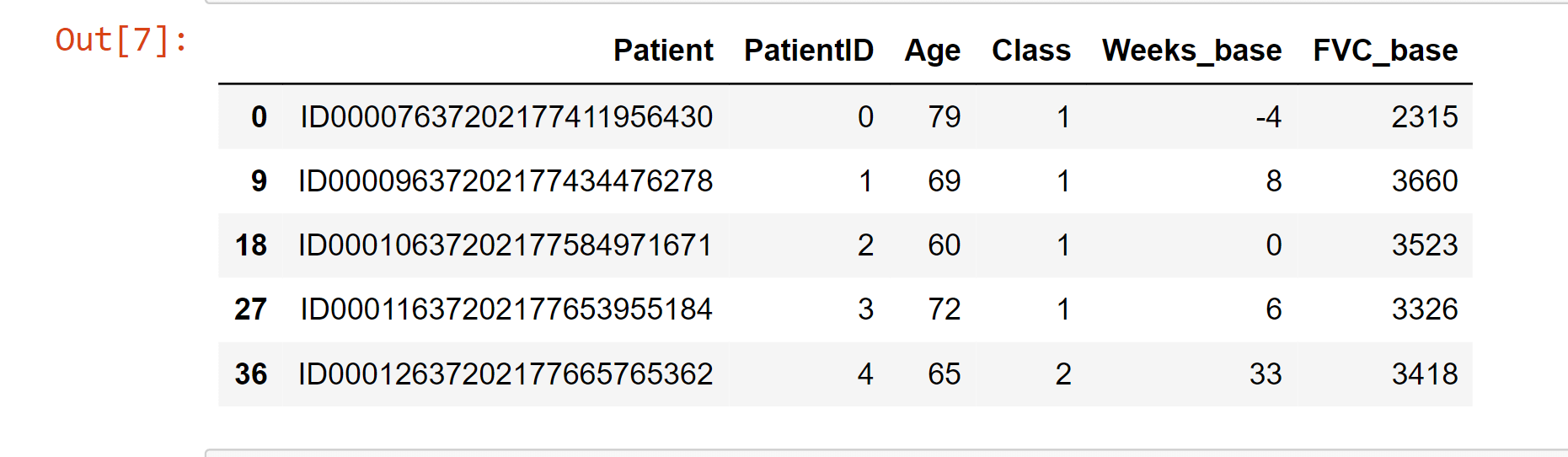
Output: 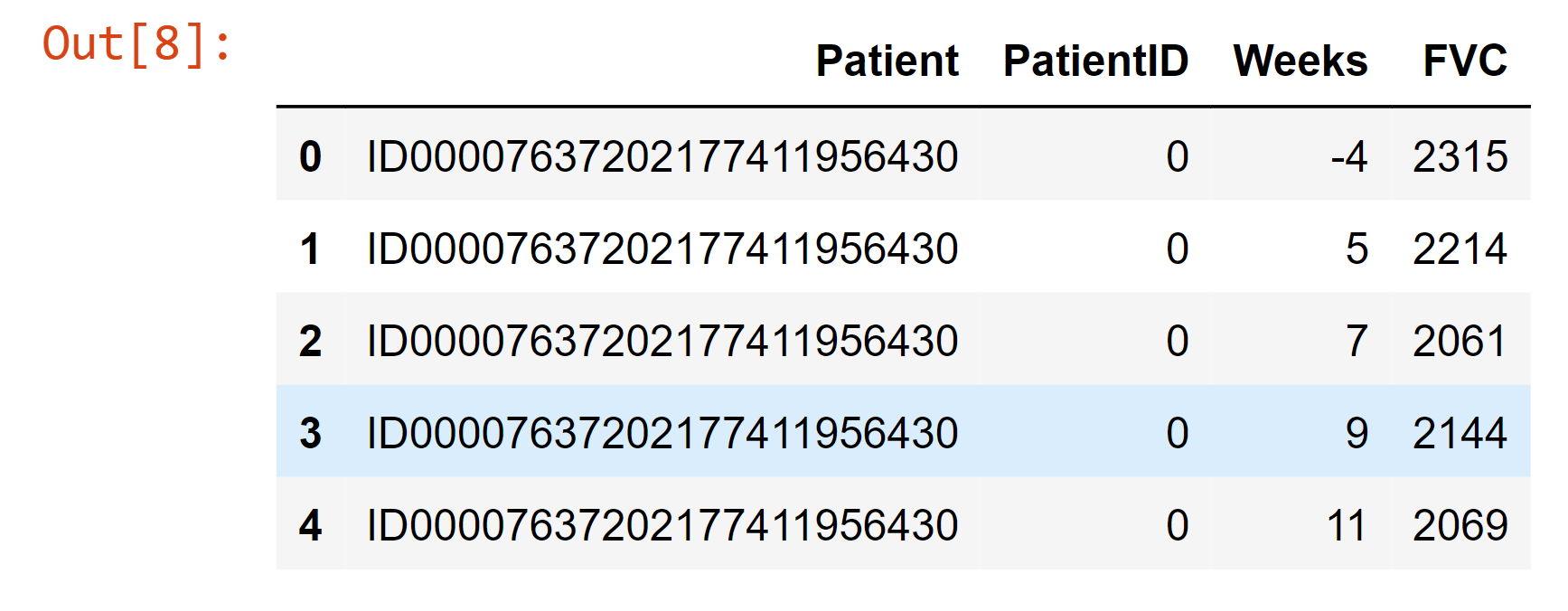
Modeling in PyMC3Fit the modelOutput: 
We just sampled 4000 distinct models that account for the data. Check the modelLet's have a look at the generative model we developed. Output: 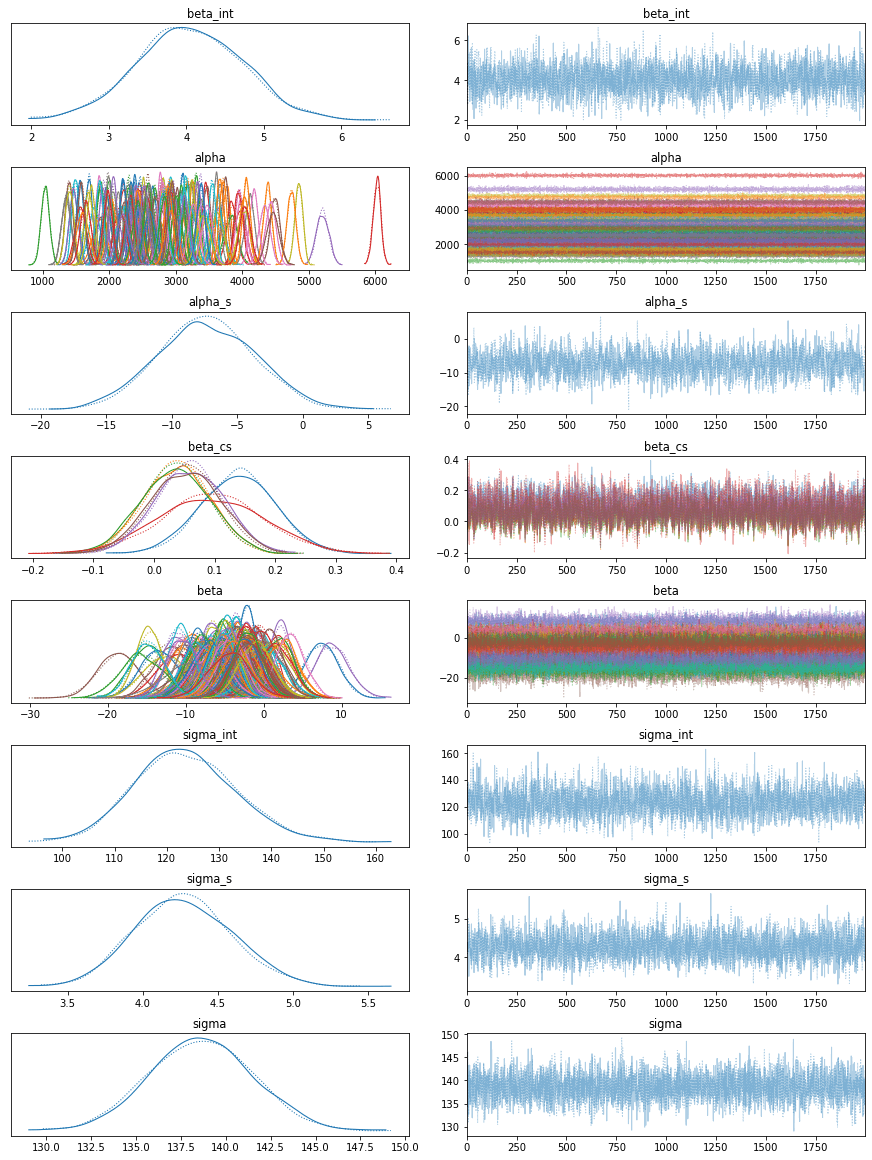
It appears that our model has learned unique alphas and betas for each patient. Checking some patientsArviZ, an extremely potent visualization tool, is included with PyMC3. Nonetheless, we make use of Seaborn and Matplotlib. Output: 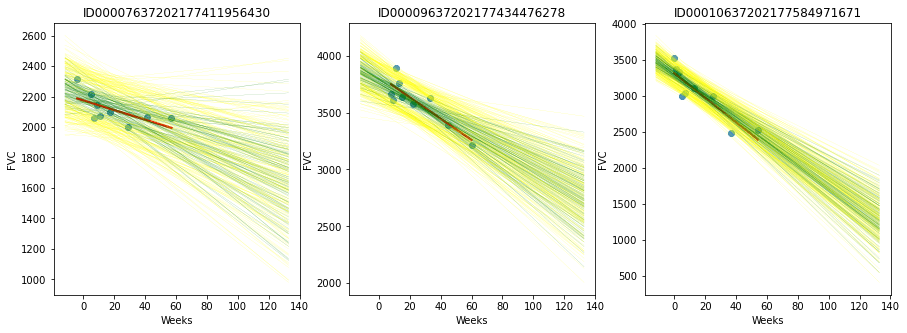
100 of the 4000 unique models that each patient possesses is plotted here. The fitted regression line is shown in green, while the standard deviation is shown in yellow. Let's put it all together! (Iterate and) Use the modelLet's use our generative model now. Simple Data Pre-processingOutput: 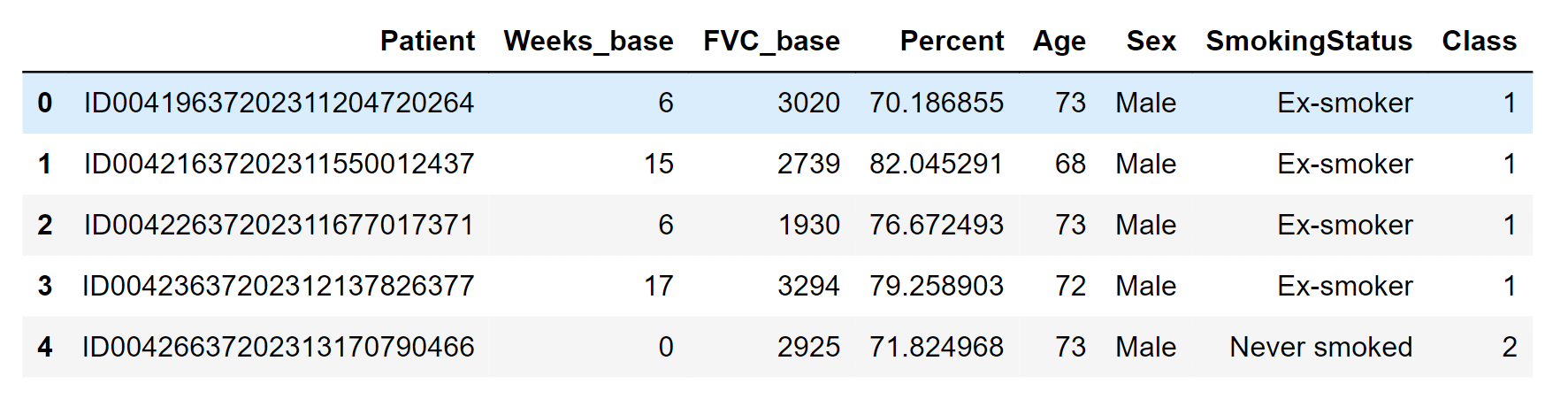
Output: 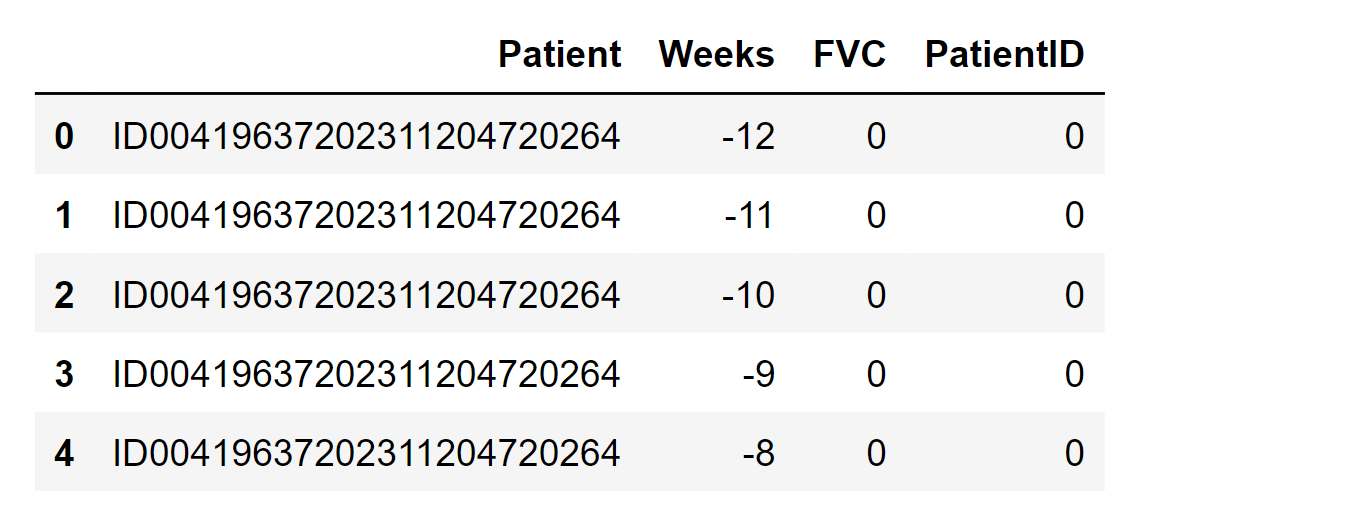
Posterior PredictionPyMC3 offers two methods for making predictions on held-out data that has not yet been viewed. Using theano.shared variables is part of the initial step. We only need to write 4-5 lines of code to complete it. We tested it, and while it did work flawlessly, we will also utilise the second strategy for greater comprehension. Although it's a tiny bit longer than the 4-5 lines of code, We find it to be far more instructive. Developers of PyMC3 explain the concept in this response from Luciano Paz. Using the distributions for the parameters learnt on the first model as priors, we will build a second model to predict FVCs on hold-out data. We continuously update our models in accordance with the Bayesian methodology as we gather new data. Output: 
Output: 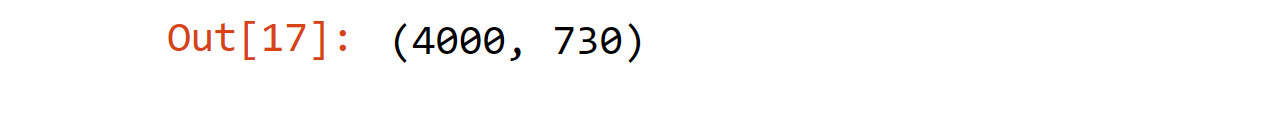
Let's go! 4000 forecasts for every point! Generating Final PredictionsOutput: 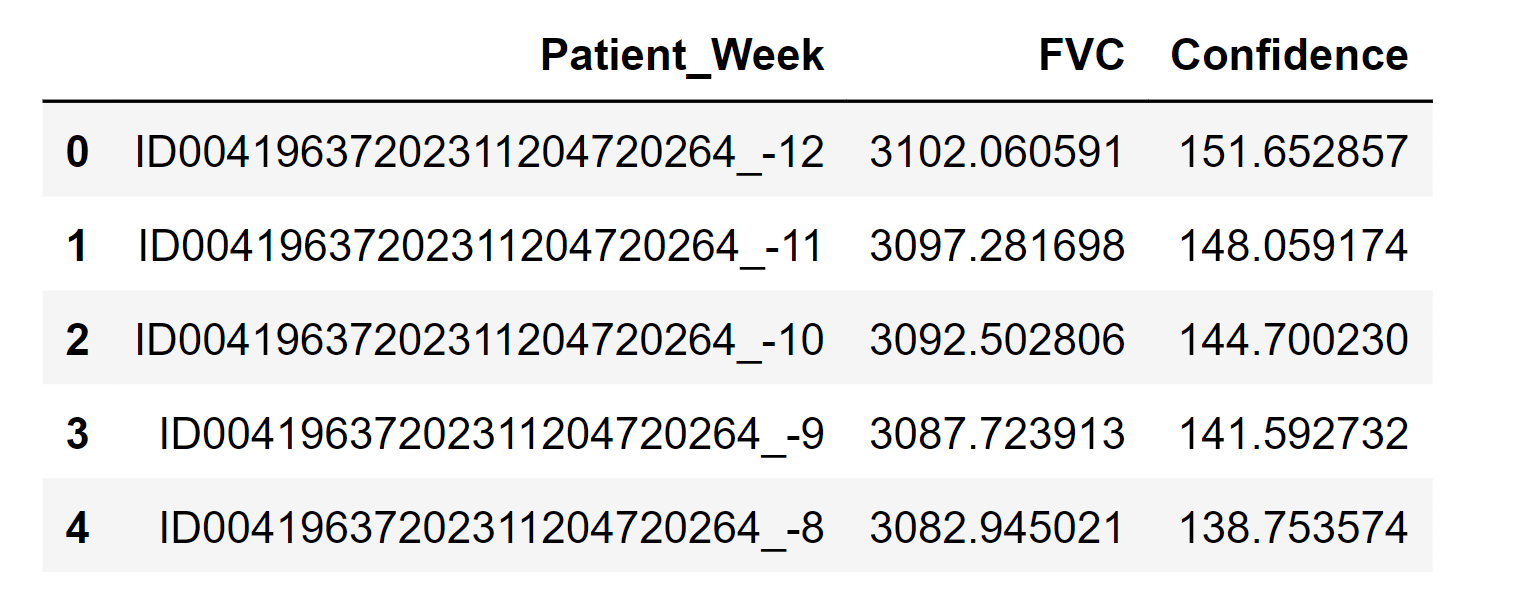
Note: We generate the final prediction so that we can submit it to the competition for evaluation.ConclusionAt its core, a probabilistic model is simply a model that incorporates uncertainty. In machine learning, this often involves representing the relationships between different variables in a system using probability distributions. For example, in a classification task, a probabilistic model might represent the probability of a particular input belonging to each possible class.
Next TopicSurvival Analysis Using Machine Learning
|
 For Videos Join Our Youtube Channel: Join Now
For Videos Join Our Youtube Channel: Join Now
Feedback
- Send your Feedback to [email protected]
Help Others, Please Share









