Sequence Classification- Machine Learning
Sequence classification is a type of problem in machine learning where the input data is a sequence of data points, and the goal is to predict a class label or a category for the entire sequence. In sequence classification, the model is trained on a labeled dataset of sequences and their corresponding class labels. The model typically consists of several layers of neural networks, such as recurrent neural networks (RNNs) or long short-term memory (LSTM) networks, which can capture the temporal dependencies and patterns in the sequence. The input data is preprocessed into a numerical form, such as a sequence of vectors, before being fed into the model. The output of the model is a probability distribution over the possible class labels, and the predicted label is typically the one with the highest probability. Sequence classification is a challenging problem due to the variable length of sequences, the presence of noise and variability in the data, and the need to capture long-term dependencies and patterns in the sequence. Nonetheless, it has important applications in various fields, including natural language processing, speech recognition, and bioinformatics. Examples of sequence classification problems include:
Python ImpleteationNow, We are going to classify Protein sequences that are HBPs or NON-HBPS for the sake of implementation. 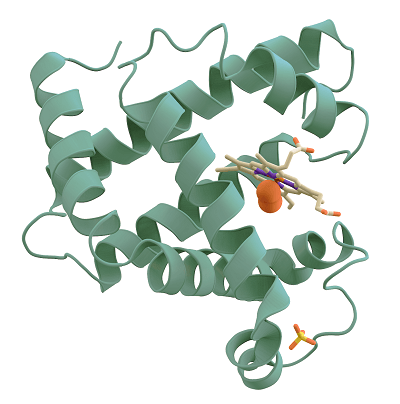
Output: 
EDA stands for Exploratory Data Analysis. It is an approach to analyzing data in order to summarize its main characteristics and gain insights into its underlying patterns and structure.
HBP (Histidine-rich basic protein) is a protein found in seminal mammalian plasma that is involved in the fertilization process. It is a small, highly basic protein that contains multiple histidine residues and is thought to be involved in the regulation of sperm motility and capacitation. In protein sequences, the term "HBP" can also refer to other proteins or protein domains that contain histidine residues and have a similar basic charge. These proteins may be involved in a variety of biological processes, such as DNA binding, transcriptional regulation, and enzyme activity. Here we will take a look at the HBP dataset. Output: 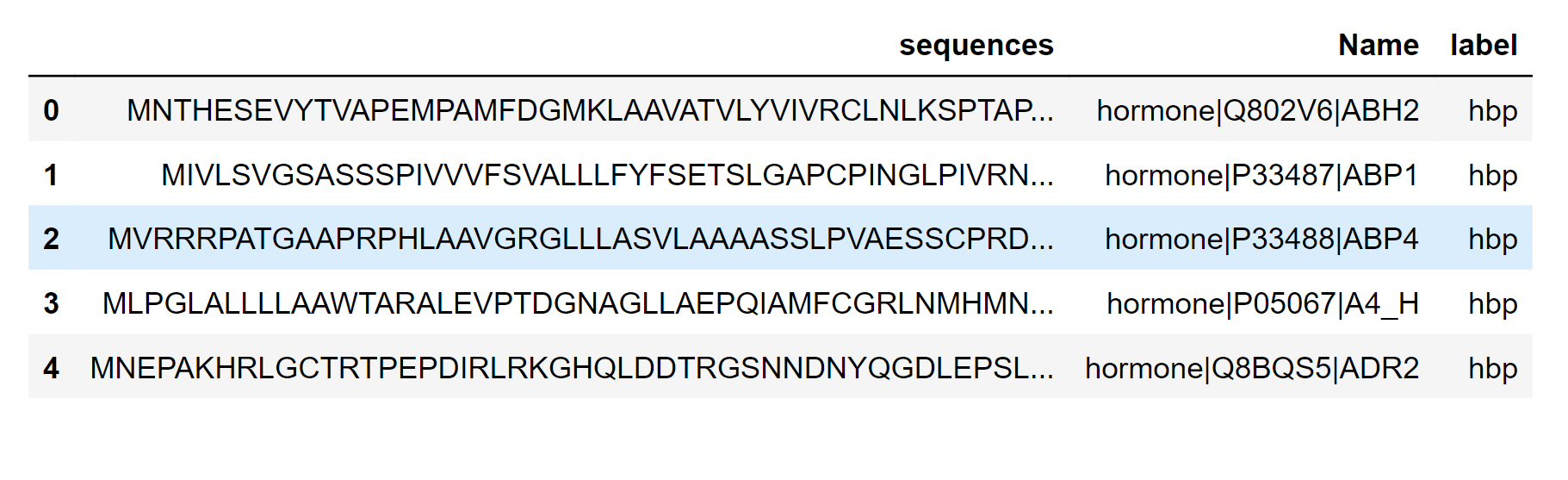
In protein sequences, the term "non-HBP" refers to proteins that do not contain a high proportion of histidine residues and do not have the same basic charge as HBP proteins. Non-HBP proteins can be involved in a wide range of biological processes, including enzyme activity, signal transduction, structural support, and DNA binding and regulation. Here we will take a look at the Non-HBP dataset. Output: 
Now we will merge both datasets so that we can create a common dataset that will contain both protein sequences. Output: 
We will use the keras library for the text processing as:
Output: 
Conv1D is a particular kind of deep learning layer for convolutional neural networks (CNNs) that is used to analyze one-dimensional data sequences, including time series, audio signals, and natural language processing (NLP) inputs. In a Conv1D layer, the input sequence is processed through a number of filters or kernels, each of which extracts a particular feature or pattern from the data. The filters slide down the input sequence one element at a time and are often tiny (e.g., 3x1 or 5x1); each convolution operation produces a single output value at each place along the sequence. Now we will train the CNN and check its accuracy on the Conv1D layer. Output: 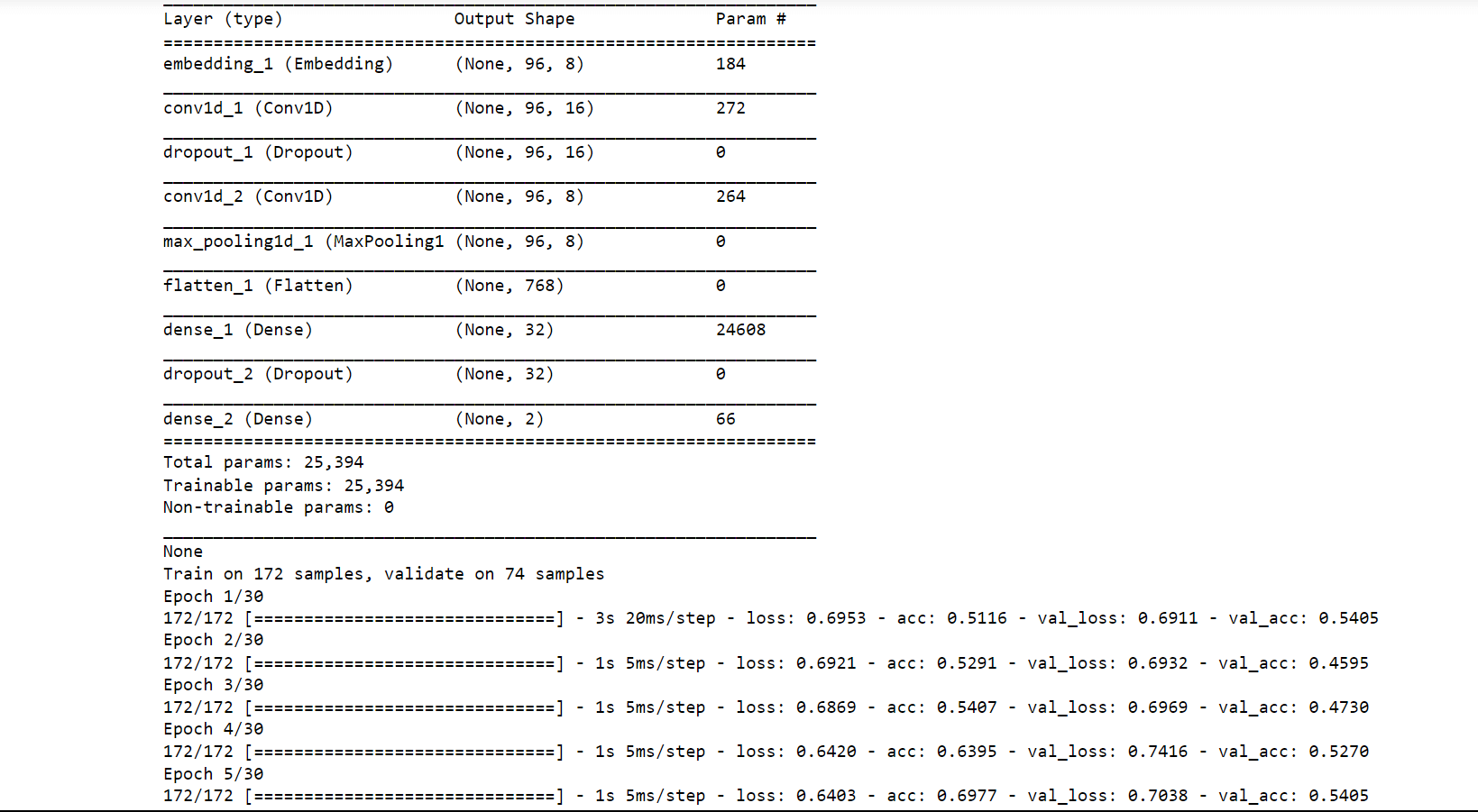
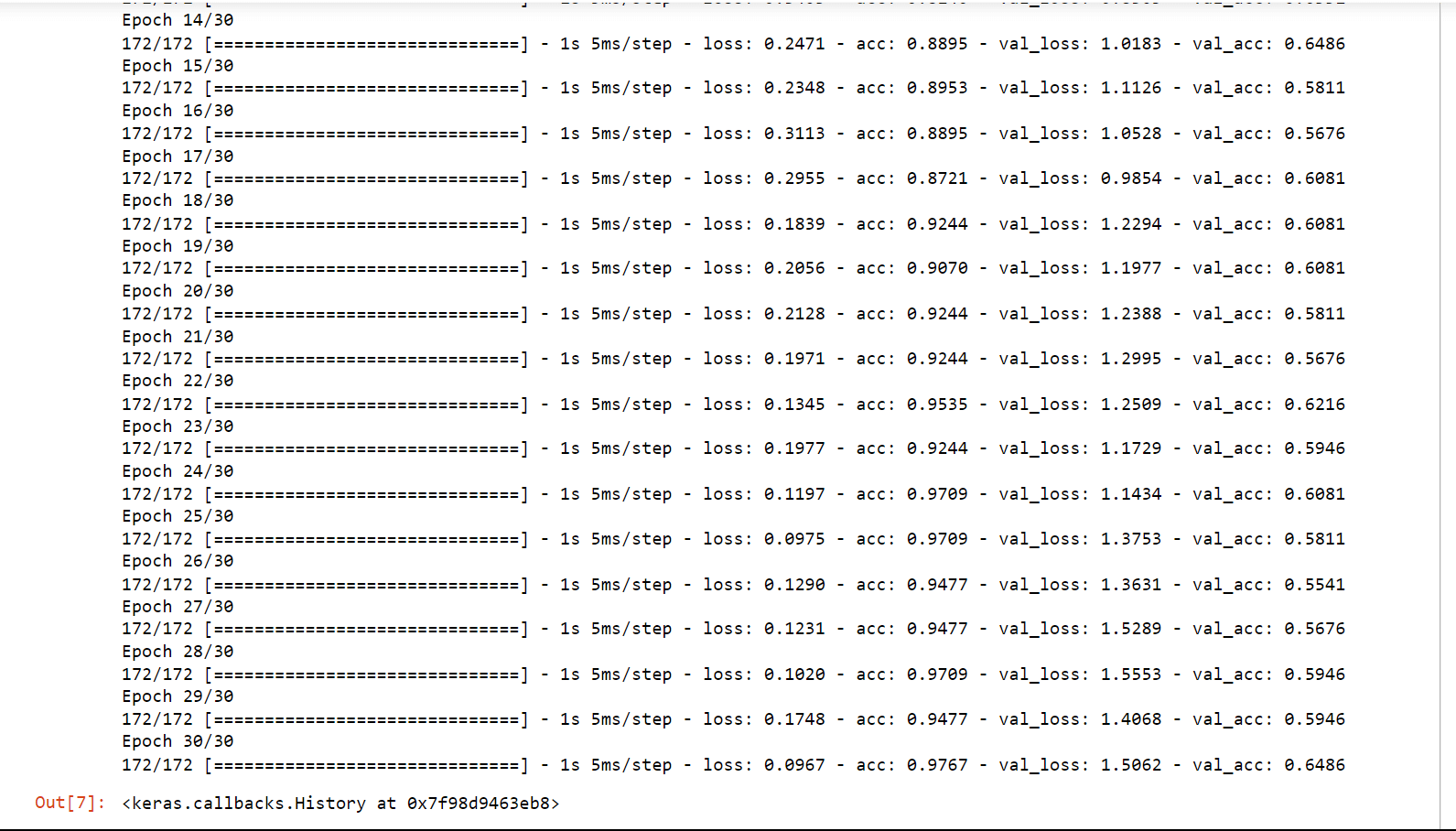
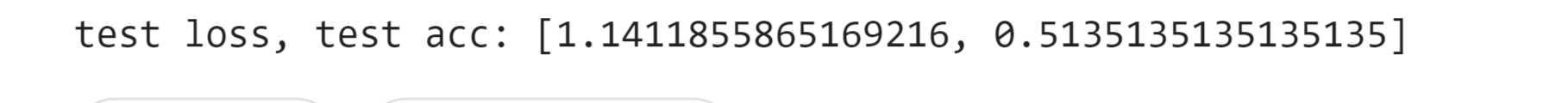
Test loss: 1.14 Test Accuracy: 0.51 We got an accuracy of 51% which is not bad considering it is quite hard to classify protein sequencing.
One-hot encoding is a process of representing categorical variables as binary vectors. This is useful when working with algorithms that cannot handle categorical data directly, such as most machine learning algorithms. We will try to encode the protein sequence into binary vectors to improve the results. Output: 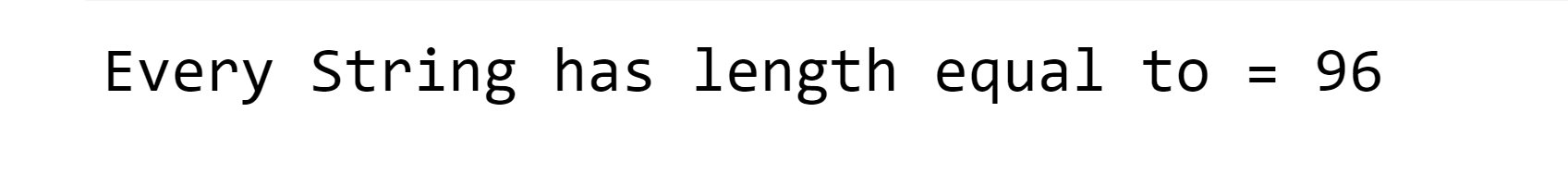
Output: 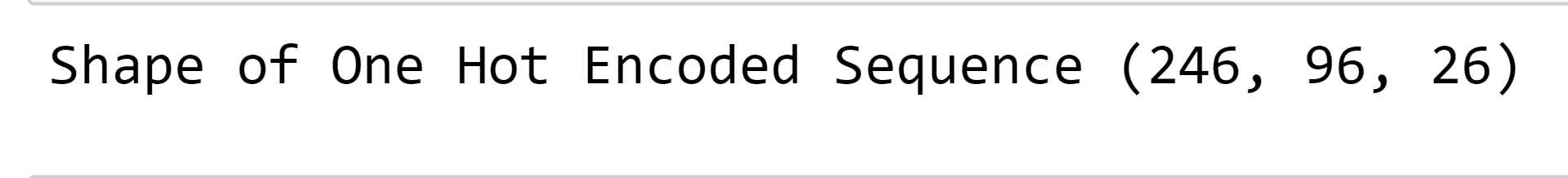
Output: 
Conv2D is a built-in layer that can be added to a model, and it is implemented using 2D convolution operation. The layer performs a convolution operation on the input image or video data using a set of learnable filters. The filters slide over the input data and perform dot products with local patches of the input to extract features that are relevant to the downstream task. Now we will train the CNN and check its accuracy on the Conv2D layer. Output: 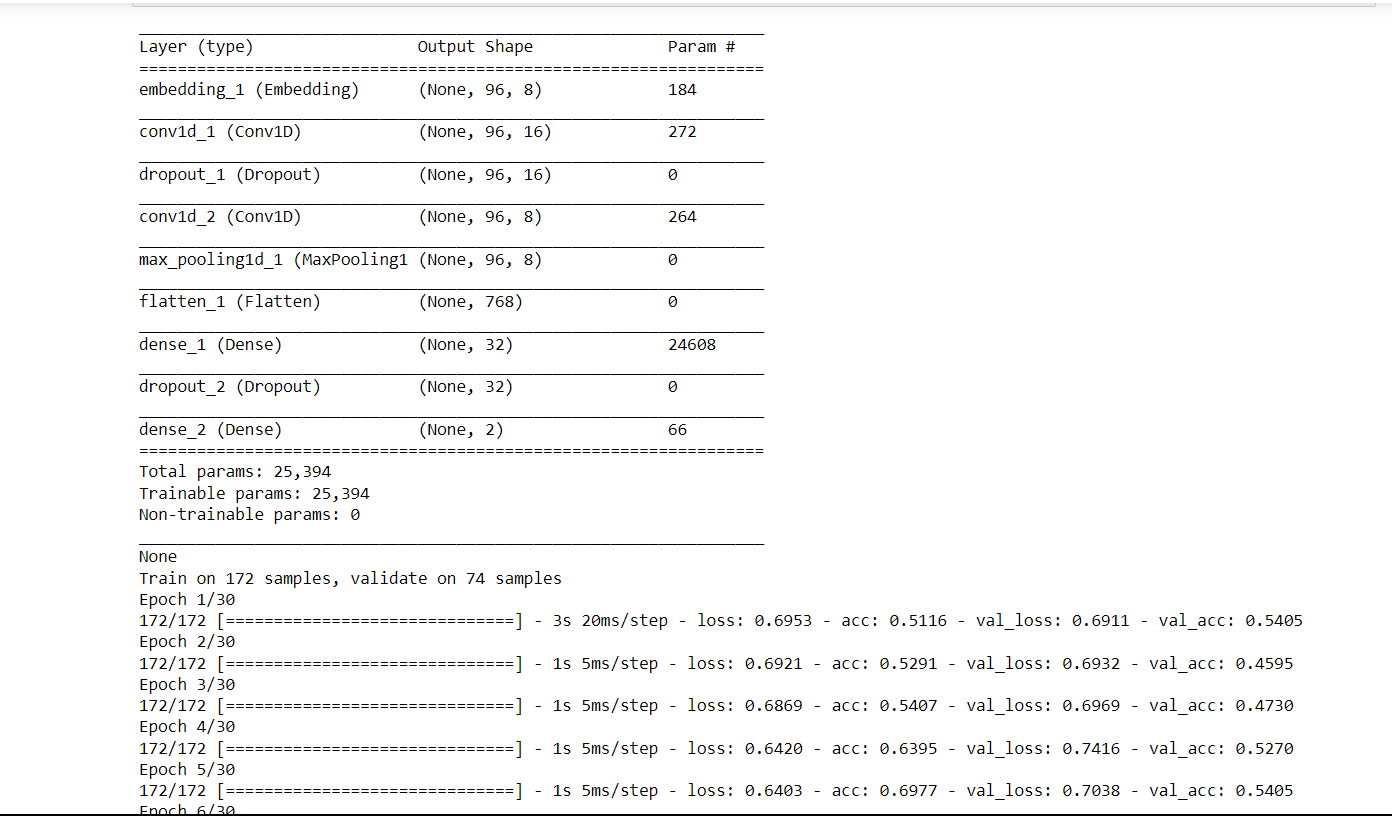
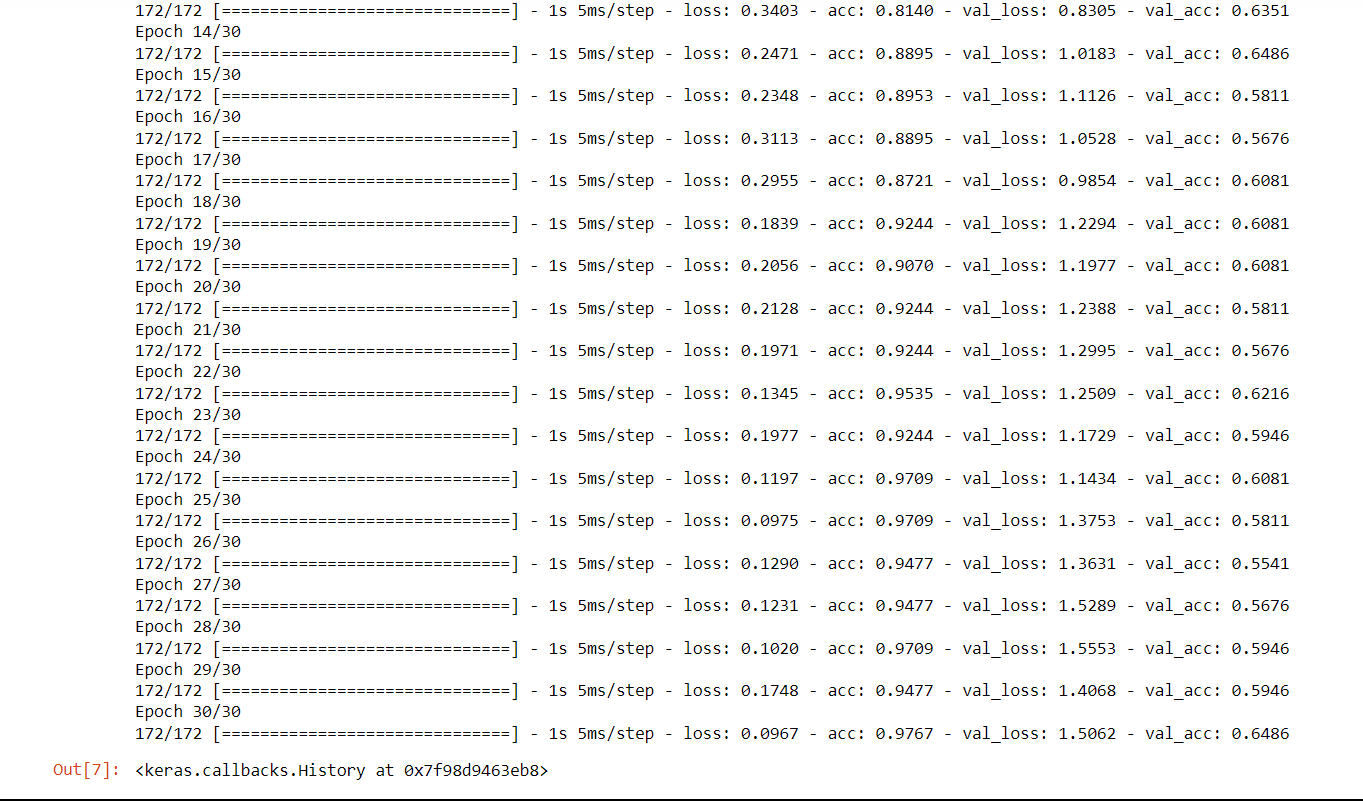

Test loss: 4.33 Test Accuracy: 0.49 Using Conv2D seems to be a bad idea for protein sequences, even though there is a huge error rate. Siamese Neural Network (Clustering ALgorithm)It is a type of neural network architecture used for learning the similarity between two inputs. It consists of two identical neural networks, called twins, that share the same weights and architectures. Its architecture is typically used in problems where the goal is to learn a similarity metric between two input samples. Examples of such problems include image or face recognition, signature verification, and text matching. Output: 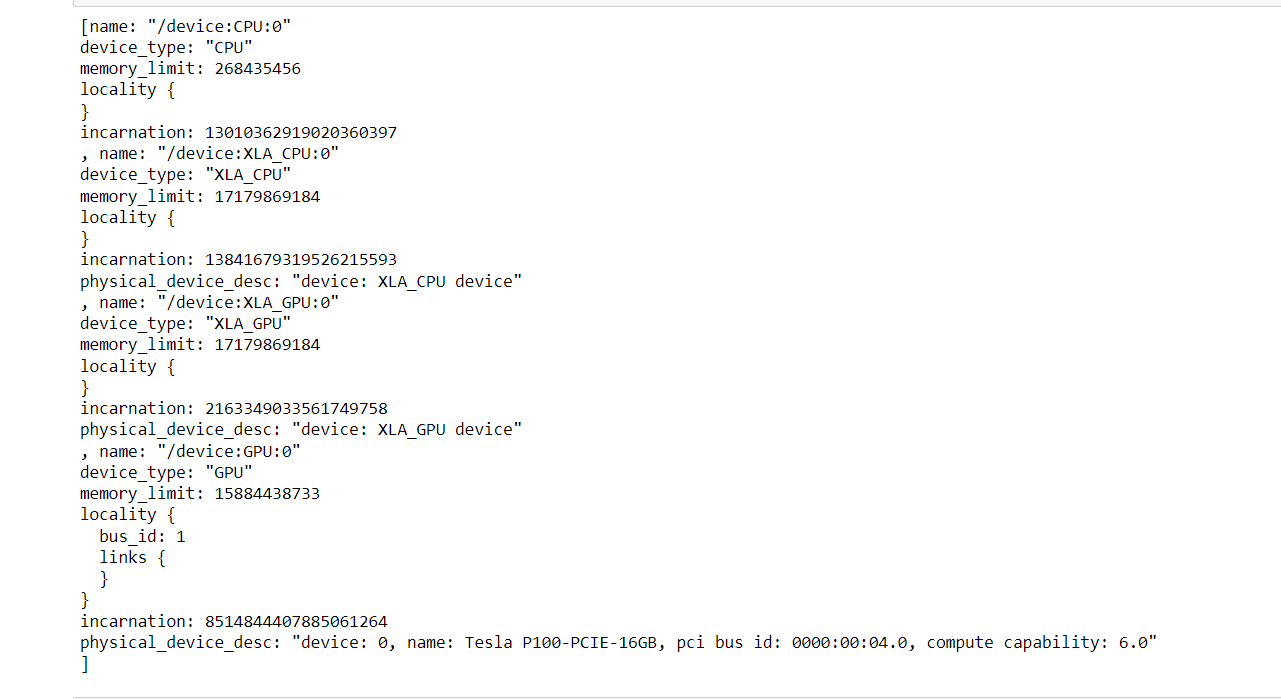
Output: 
Output: 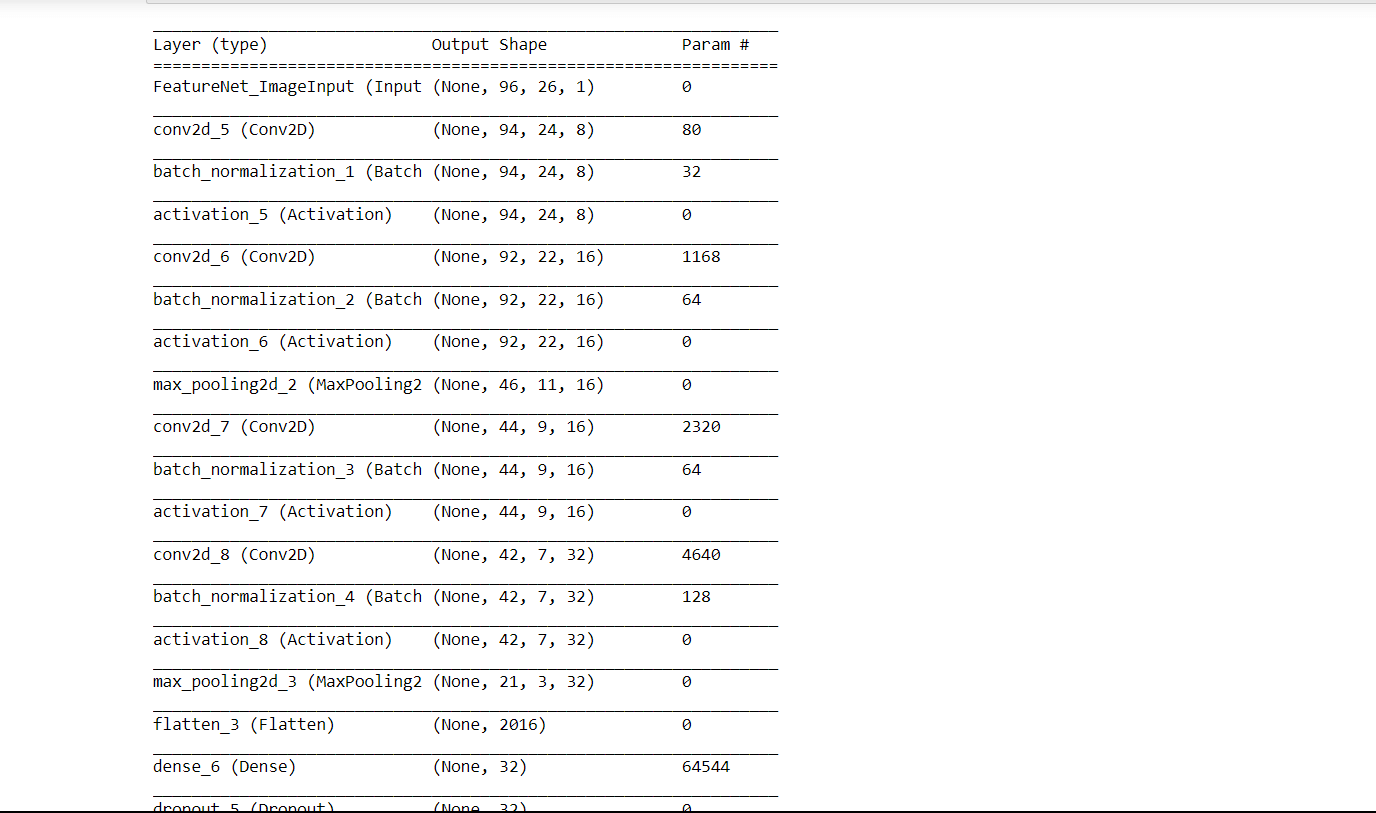
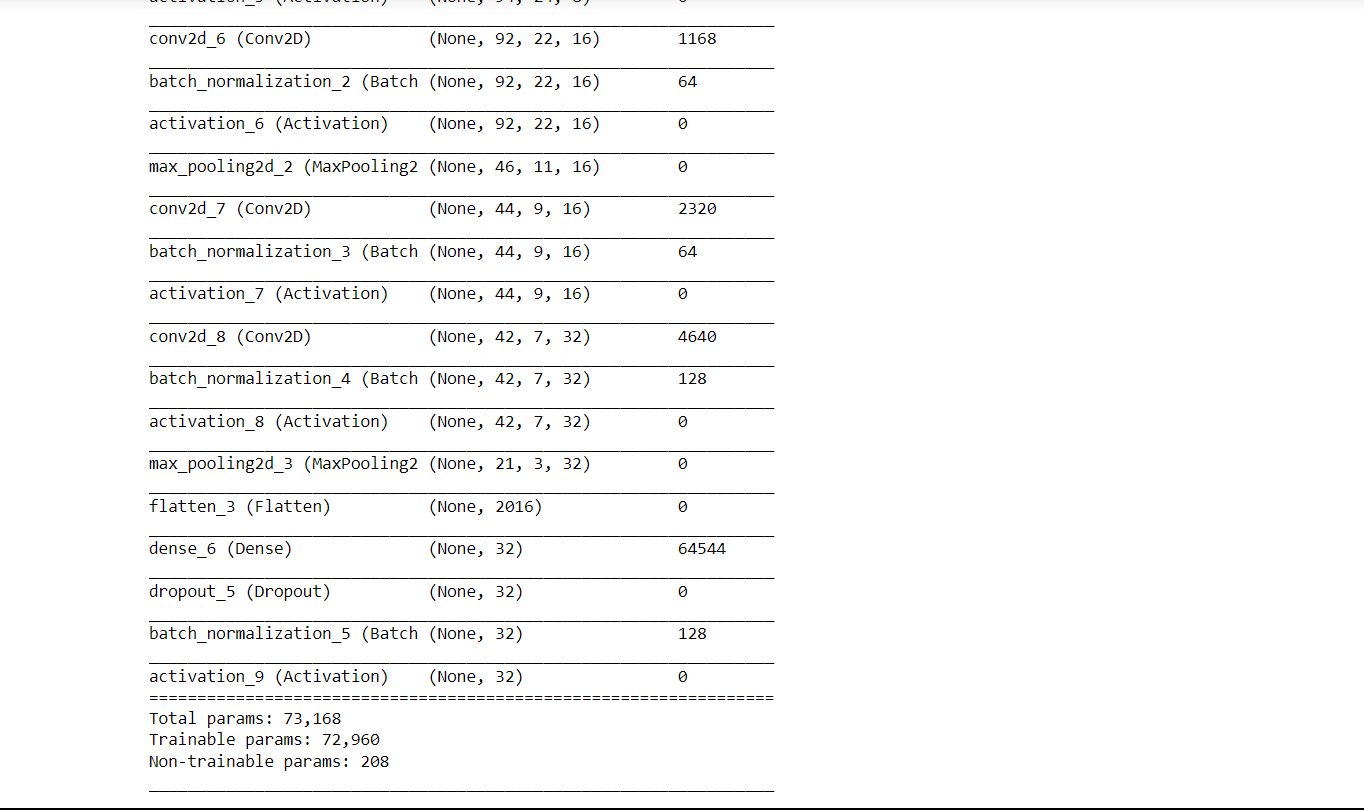
Output: 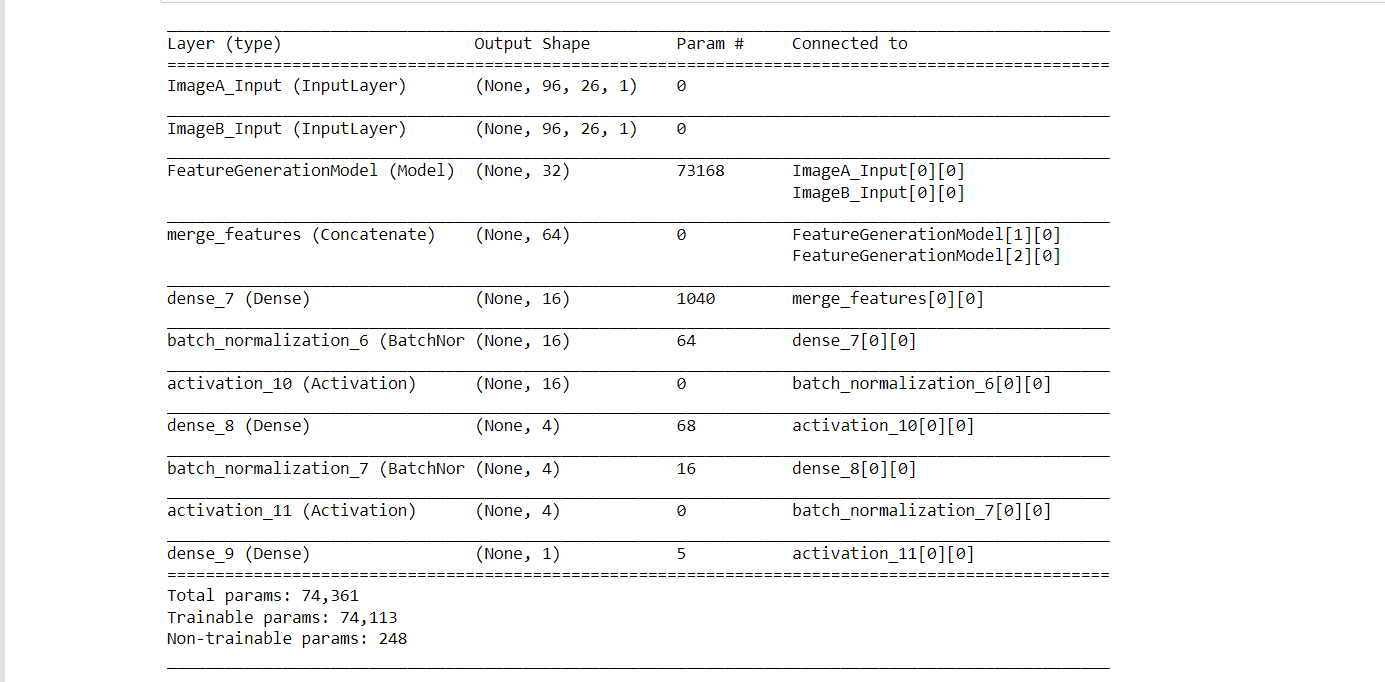
Output: 
Bag Of Words as Feature ExtractorIn the BoW model, each document is represented as a vector of word frequencies, where each dimension represents a unique word in the document corpus, and the value in each dimension represents the frequency of that word in the document. Feature Extractor is a method used to extract relevant features from textual data. We will use bags of words to extract relevant features from the data by pattern recognition. Output: 
Output: 
Other Models Here, We will classify the protein sequence with the help of other models. Output: 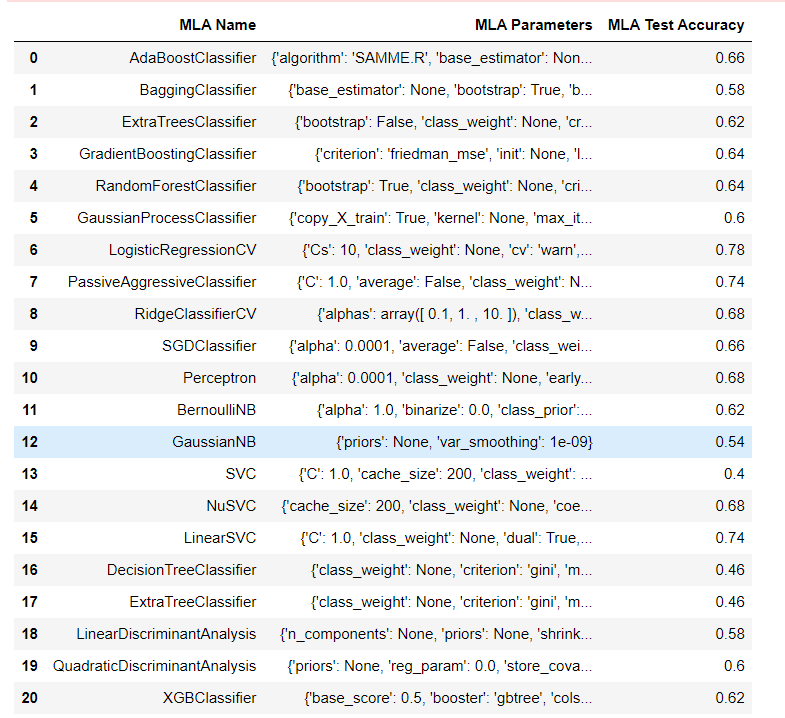
Surprisingly Logistic Regression has the highest accuracy rate of 78%, and Support Vector Classifier has the lowest accuracy rate, i.e., 40% From this, we can say that Using Logistic Regression for Sequence Classification is one of the best choices in the case of Protein Sequence Classification. ConclusionSequence Classification is an important machine learning task that has applications in a wide range of fields. Various models can be used for sequence classification, including RNNs, CNNs, and transformers. These models are trained using labeled data and can be used to predict the class or label of new input sequences. The choice of model depends on the nature of the input sequence and the specific requirements of the application.
Next Topic#
|
 For Videos Join Our Youtube Channel: Join Now
For Videos Join Our Youtube Channel: Join Now
Feedback
- Send your Feedback to [email protected]
Help Others, Please Share









